SPLITS (Spline Analysis of Time Series) is a software package to analyze time series of remotely sensed parameters such as vegetation indices. It has been designed for handling large datasets on Linux and UNIX platforms. Initially developed during my PhD thesis, SPLITS has been constantly developed and refined. The various spline models available in SPLITS can handle irregular time series by using time stamped data available now in more and more archives. MODIS vegetation index products, e.g. are time stamped starting from collection 5.
The user interface of SPLITS follows a classical UNIX toolbox concept: There is one program for every purpose, and several programs may be combined using shell scripts to manage more complex tasks. With only a few exceptions, all programs are command line based. Of course, there are programs with graphical interfaces to explore your time series imagery and try different spline models on your data before you settle for a model that you want to apply to your entire dataset using the command line tools. The command line builder integrated into the splview graphical user interface can help you to translate your graphical settings into actual commands.
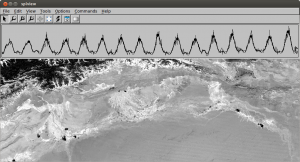 The typical workflow with SPLITS is to first prepare a time series dataset, using either the preprocessing tool (prepro) that comes with SPLITS, or your own tools. The time series dataset can then be viewed and explored using splview to select an appropriate model. To apply the model to your entire dataset, use splfit. The result is an intermediate dataset describing the fitted spline model. Finally, there exist a number of analysis programs to make use of the model, e.g. phencal to calculate phenological parameters or splcal, if you simply want to smooth or interpolate time series. Please see the manual pages and example shell scripts that come with the package for more information.
The typical workflow with SPLITS is to first prepare a time series dataset, using either the preprocessing tool (prepro) that comes with SPLITS, or your own tools. The time series dataset can then be viewed and explored using splview to select an appropriate model. To apply the model to your entire dataset, use splfit. The result is an intermediate dataset describing the fitted spline model. Finally, there exist a number of analysis programs to make use of the model, e.g. phencal to calculate phenological parameters or splcal, if you simply want to smooth or interpolate time series. Please see the manual pages and example shell scripts that come with the package for more information.
The following versions of SPLITS are available for download:
- Latest
- 1.11
- Resolved memory management and storage problems in C++ API related to phenological parameters
- Proper handling of No Data Values in splview
- Minor multithreading issues fixed
- 1.10
- Fixed some issues when compiling with gcc 7.3.0 and newer
- 1.9
- Configuration files to save and restore basic GUI settings in splview.
- Support for non-uniform knot vectors
- 1.8
- Now includes a C/C++ API for custom spline fitting and phenological calculations using plain C/C++. API headers and a static library are installed by default. If you don’t wish to use it, you can ./configure SPLITS with the –disable-api flag set.
- 1.7
- 1.6
SPLITS comes as a pretty standard Linux/UNIX installation package. To compile and install, unpack the package tarball into a directory of your choice and make it the working directory, e.g. by the following commands:
tar -zxf splits-1.10.tar.gz cd splits-1.10
Many of the components required by SPLITS are most likely already installed on an average Linux or UNIX system. However, you may have to install the following additional libraries: GDAL for reading and writing image data, Armadillo for numerical computation and optionally FLTK for graphical user interaction. Using FLTK can be disabled on systems without a graphical user interface (e.g. servers), see below. To compile the source code and install the resulting binaries to the default location, type something like:
./configure make sudo make install
It is likely that you need to tweak certain configuration options on Linux or UNIX systems of different flavors. On Ubuntu, I usually configure with
./configure CPPFLAGS="-I /usr/include/gdal"
In server environments, you might want to use SPLITS without the graphical user interface. The following option suppresses the installation of SPLITS programs with graphical interfaces:
./configure --disable-gui
The following command will print all available configuration options:
./configure --help
After successful installation, type
man splits
to get started. If you prefer printed documentation rather than manual pages, you can build Postscript or PDF versions of the manual pages:
make pdf make install-pdf
The install command will copy the PDF files to a location where your system stores documentation (most likely /usr/local/share/splits/). The documentation directory also contains examples on how you can use the software. SPLITS can be used under the terms of the GNU General Public License.
